
Here’s a brief look at the discussion of aDNA in Denisovan Origins and what Collins and Little got wrong.
In their book, Denisovan Origins, authors Andrew Collins and Greg Little appear to paint a picture of how giant Solutreans of “Denisovan origin” dominated less capable Native Americans of a more diminutive stature who arrived across the Beringian land bridge: showed them how to build mounds, established themselves as their “elite” leadership, and passed on their “advanced knowledge.” You know… because plain old Indians couldn’t possibly have figured out all the cool stuff.
The very notion is racist. Are Collins and Little racist? Probably not intentionally. But not all racists are card-carrying members of the KKK or NAZI’s or any number of other extreme notions you can think of. Simply not willing to afford an indigenous population the credit its due for its own accomplishments without injecting some silly notion of “elites” and “giants” is racist.
But perhaps it’s a racist notion arrived at innocently through fallacious thinking. Here are the problems with Collins’ and Little’s understandings of ancient DNA (aDNA).
Haplogroups X, X2, and X2a
In his section of the book, Little repeatedly states that Haplogroup X exists in the aDNA of Native American populations (chapters 23 and 26). Occasionally he rightly refers to the Native American haplogroup in question as X2a but sometimes he calls it X2.
These are three different haplogroups.
I’ll put some quotes from Collins and Little below followed by a critique:
Haplogroup X also is more problematical. While haplogroup X is present in two versions (X2 and X2a) in about 3 percent of living Native American tribal members, it is not evenly distributed.”
Collins and Little: Denisovan Origins, Chapter 26
The only thing he really got right here is the last phrase of that sentence. Distribution is not even. The two X haplogroups found in ancient Native Americans are X2a and X2g. The latter is rare, but both are uniquely North American. They don’t show up in the aDNA of other populations in the world. Nor do other X haplogroups show up in the aDNA of North American populations, at least not in current data (it could all change in the future with better aDNA surveys of ancient populations. Regardless, there is no X2 or X in the current available data for ancient North American populations. Little is wrong about this.
“The X2 haplogroup found in the Altai Mountain regions in Siberia closely matches the X2 found in Native Americans. Of course, many archaeologists hailed this as proof that X2 also came into the Americas via Beringia.”
Collins and Little: Denisovan Origins, Chapter 26
The haplogroup found among populations in the Altai Mountain region of Siberia is X2. But I’m not aware of any archaeologists that “hail this as proof” that the X2a haplogroup found in North America came via Beringia. Other lines of evidence are suggestive of this, however, which David Madsen (2015) lists in nice detail in the very reference Little cites for this next sentence.
However, the X2a version has also been found in heavy concentration in the Orkney Islands of Scotland.
Collins and Little: Denisovan Origins, Chapter 26
For this he cites David Madsen (2015). First, there is no X2a haplogroup on Orkney. Not in among ancient populations and probably not among modern ones. There is, however, the X2 haplogroup present on Orkney Island. And this is what Madsen says: “although, as an aside, it is interesting that a modern population with one of the highest percentages of the X2 clade, higher even than Native American populations, is found in the Orkney Islands off the coast of Scotland.”
It’s important to note that Madsen isn’t saying that X2a is on Orkney. Nor is he saying that X2 is among Native Americans. He’s saying that the percentage of Orkney’s X2 is higher than the percentage of Native American X2a. If you’re not reading the entire paper carefully and if you don’t already have a very basic, working understanding of aDNA (either nuclear or mitochondrial), you might miss the context. But Madsen is assuming his readers know X, X2, X2a, and others are different haplogroups. And that X2a (along with X2g, X2j, X2b, X2c, …) is in the X2 clade.
The real question is this: is Little truly ignorant of what a haplogroup is and isn’t? Or is he intentionally conflating and confusing them for his readers. My suspicion is the latter. It’s useful for for the conclusions he and Collins began with when they started looking for data. By pseudoscientifically reducing the X2a haplogroup to the X haplogroup, they can fit data into the very conclusions they’ve both carried for years: giants dominated Native Americans, etc.
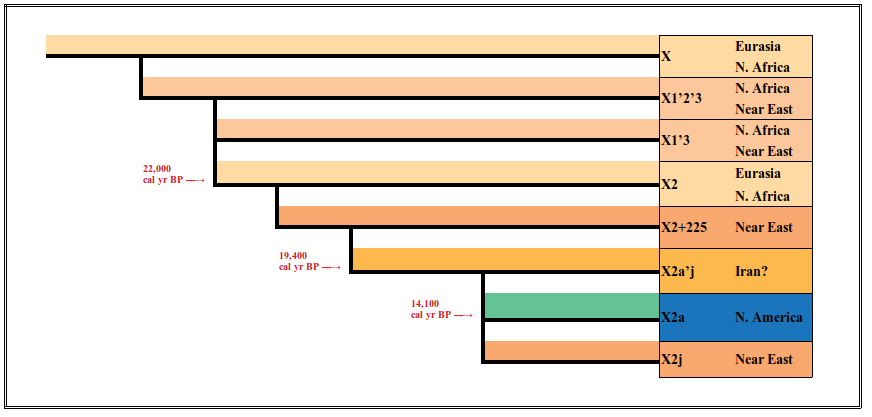
Maybe X2a did come to North America by way of an Atlantic crossing. The Solutrean hypothesis is science. But it’s an hypothesis. To date, the evidence stacks against it. And haplogroup X isn’t its savior because that haplogroup is not present in North America. Nor is hapogroup X2 (thehaplogroup found in a modern Orkney population). The haplogroup X2a lineage doesn’t simply fall from X to X2 to X2a. The X2a’j clade lies between X2a and X2-225 before stepping back to X2, along with nearly 10,000 years of mutations.
There’s no good reason to think an Atlantic arrival for X2a is more likely than the Beringia route. Little liked to suggest that since X2a was observed in North America’s northeast that this is evidence, but it really isn’t. X2a was observed in Kennewick man (8690-8400 cal yr BP), putting it about as early as any X2a samples. Little tries to cover this base:
Kennewick Man is one of the few haplogroup X skeletons found that far Northwest. Perhaps the greatest importance of Kennewick Man is that his lineage is haplogroup X2a. It is proof that X2a was in the Americas long, long ago, and not that he was from Asia.
Collins and Little: Denisovan Origins, Chapter 26
At least Little acknowledges that the haplogroup is X2a and not X2 or X. But he’s dead wrong when he says its proof Kennewick was not from Asia. In fact, Kennewick showed many Siberian genetic affinities and no recent European ancestry.
Little isn’t the only one of the two authors to get all this wrong. Here’s Collins in chapter 10:
The closest source of haplogroup X outside of the northeastern and eastern coasts of North America is that present among modern populations in southwestern Europe, which just happens to be the very same territory in which the Solutreans thrived circa 20,000–15,000 BCE.
Collins and Little: Denisovan Origins, Chapter 10
He is, of course, wrong. I’ve included a chart based on the one found in Raff and Bolnick (2015) to give you an idea. But suffice to say, Haplogroup X and its sub-clades are present in more places than Collins seems aware. Moreover, he’s also conflating haplogroup X with X2a. And they both occasionally conflate modern populations with ancient populations. As if the presence of haplogroups in modernity reflects the localities of antiquity. They both seem to have an affinity with the X2 haplogroup in modern Orkney populations which 1) is irrelevant to the X2a haplogroup in the Americas; and 2) representative of modern populations, not those of 14,000+ years ago.
Here’s Collins again in chapter 16:
The tribe possessing the highest level of haplogroup X is the Ojibwa. Up to 26 percent of its population’s mtDNA contains haplogroup X, while the Cree also possess haplogroup X, but, once again, at a slightly lower level than their southern neighbors, the Ojibwa.
Collins and Little: Denisovan Origins, Chapter 16
For this, Collins cites Brown et al. (1998), referencing Table 3 on page 1858. You might be wondering why Collins and Little are citing a 20+ year-old paper when there are many more up-to-date sources. The reason, of course, is that within just a few years of 1998, sub-clades of X were being identified and understood. Collins and Little would have been better off citing Fagundes et al. (2008) in the very same journal. But the obfuscation of haplogroup X and ignoring the implications of how the sub-clade X2a is defined, makes it easier for Collins and Little to fit their narrative into the pre-conceived conclusion they already have.
Writing of haplogroup X in broad terms in a book that deals with the peopling of the Americas isn’t really wrong. Unless you’re trying to ignore the nuances and details that come along with its sub-clades. X2 in Orkney is a “modern population” (Madsen 2015: 213). X2a in North America is without a “clear record” of its evolutionary history “in any population” (Raff and Bolnick 2015: 298; see also Fernandes et al. 2012).
Mystery Mongers
Wherever there’s a mystery in science, there will probably be someone ready and willing to pick it up and repackage it as proof of whatever silliness they’ve already concluded to be true. In this case, Collins and Little (among others, eg. Graham Hancock, Brien Foerster, etc) have quickly picked up on the fact that very little is known about Denisovans. With a handful of skeletal remains that point to individuals of large stature, they see the opportunity to display this as proof of their giants. With the mystery of the origin of the X2a haplogroup, this is now an opportunity to point to Solutreans, who are important for Collins and Little to get their giants across the Atlantic and into the Americas.
Never mind that there are literally only a handful of skeletal remains for the Denisovans: a piece of a finger, a tooth, a portion of a mandible, a few others… perhaps as many as seven pieces in all. You needn’t be a bioarchaeologist to realize only the more robust bones of the most robust individuals in the best of conditions will survive the tens of thousands of years these tiny fragments of even fewer individuals. It would be no surprise to me if, like my family, the Denisovan analog these remains are from included some very tall individuals and some rather small members.
And to suggest that because there are X2 haplogroups present in modern populations on Orkney (or even ancient populations for that matter) and therefore it’s more likely that X2a came from there is showing an extremely poor understanding of how haplogroups work, since X2a is not closely derived from X2.
References and Further Reading
Brown, Michael D., Seyed H. Hosseini, Antonio Torroni, Hans-Jürgen Bandelt, Jon C. Allen, Theodore G. Schurr, Rosaria Scozzari, Fulvio Cruciani, and Douglas C. Wallace (1998). mtDNA Haplogroup X: An Ancient Link between Europe/Western Asia and North America? The American Journal of Human Genetics, 63(6), 1852–1861.
Fagundes, Nelson J.R., Ricardo Kanitz, Roberta Eckert, Ana C.S. Valls, Mauricio R. Bogo, Francisco M. Salzano, David Glenn Smith, Wilson A. Silva Jr., Marco A. Zago, Andrea K. Ribeiro-dos-Santos, Sidney E.B. Santos, Maria Luiza Petzl-Erler, and Sandro L. Bonatto (2008). Mitochondrial Population Genomics Supports a Single Pre-Clovis Origin with a Coastal Route for the Peopling of the Americas. American Journal of Human Genetics 82: 583-592.
Raff, Jennifer A. and Deborah A. Bolnick (2015). Does Mitochondrial Haplogroup X Indicate Ancient Trans-Atlantic Migration to the Americas? A Critical Re-Evaluation. PaleoAmerica, 1(4), 297-304.
Carl Feagans paints Andrew Collins and me as asserting a racist notion in Denisovan Origins. The very first reason he cites is that we wrote that “giant Solutreans of ‘Denisovan origin’ dominated less capable Native Americans of a more diminutive stature who arrived across the Beringian landbridge: showed them how to build mounds.”
Hmm. Now I have already issued an article about one error and definite mistake I made in the book, but it’s certainly not this. To get right to the point: What Feagans asserts is either a blatant lie or for his own psychological reasons he just ignored a lot of the book.
In Denisovan Origins I stated very, very clearly that mound building in the Americas began in South America, around 4,000 years before any mounds were erected in North America. The Clovis Culture, which we assert (like many other archaeologists was a remnant of Solutreans), NEVER reached South America. From South America, mound building gradually moved North. In addition, the first pottery in the Americas was made everywhere in South America before it went North, thousands of years later. The “First Native Americans” were in South America. South American archaeologists have claimed that the old, white, male American archaeologists dominating in North America ignore and dismiss their findings and research.
We asserted, as many South American archaeologists do, that South America was inhabited via a southern route many, many thousands of years before North America. There were many migrations into N and S America (including from Beringia) before a small band of Solutrean descendants arrived around 15,000 years ago or so. And, as we asserted, the Solutreans were Asian in origin. We assert that all of the peoples in the Americas prior to historic times were Native Americans with their ancestry origin somewhere in Asia.
After the Younger Dryas event, the cultures in N, S, and Central America all developed their own characteristic and unique styles involving mounds, earthworks, pyramids, and stone structures.
With respect to the mtDNA things cited, I’ll relate what I stated in the book. There are literally thousands of genetic research articles on it, and if you search around you can find something that confirms what you believe or want to believe. And I added that a lot of it is no more than educated guesswork based on assumptions. Yes, that applies to us. But it also applies to you. As I wrote, confirmation bias relates to starting out with a belief system and then one seeks out confirming evidence while ignoring or dismissing evidence that doesn’t confirm the belief.
It is interesting that you end with a description of the very fallacy your book is replete with. It’s additionally interesting that you spend five of six paragraphs addressing that which I hardly discussed at all. And only a single paragraph regarding the errors in representation of DNA which is what the entire critique was about.
This is all part of fitting the data to suit the preconceived conclusion that Collins and Little began with before they started the book together. Little waves his hand and essentially states, “you can find any data you want to support any idea you have. We did” in that last paragraph.”
The problem with the pseudoscientific methods of data-mining is that cherry picking doesn’t really work. Little and Collins didn’t simply look for sources they liked, they obfuscated the data they used by redefining what it means to be X2a and X2. And by ignoring the scientific methods used in genetics if they were contrary to the conclusions they already had.
While the “Solutrean hypothesis” is a genuine hypothesis, it isn’t one that is, to date, supported by good evidence. There simply is no good reason as yet why the X2a haplogroup couldn’t have arrived along with everyone else across the Beringian land bridge. Nor is there any DNA evidence that supports a southern route. The aDNA signals of Austral-Asian ancestry in Brazil are more likely the result of the same land bridge route.
With regard to the article about the one definite mistake Little made, I’ve yet to come across it. But I imagine it’s the part where he’s calling mtDNA “non-human” DNA. Regardless of the origins of mitochondria (a leading hypothesis is that mitochondria evolve from bacteria), humans have and depend on mitochondria in their cells. They’re a big part of ATP production, so they’re definitely as human as any cell.
Mr Feagans another good write up, congratulations.
I have one question about the Haplogroups, do they come from both faith rand mother and do they mix?
That’s a really good question and I purposely didn’t go into the basics of DNA since that would easily be a chapter if not a book all to itself.
But very briefly, mtDNA only comes from the mother. So the X2a (or any haplogroup) would be passed down from only the maternal line. So, you have mtDNA in your body, but your children would never inherit it (though there are some very rare, hypothesized exceptions–a subject that might have *its* own book!).
The DNA that resides in the mitochondria in your body came from your mother. And that mtDNA came from her mother… and so on.
Haplogroups are derived from mutations and geneticists can apply a bit of math to observations and make some fairly good predictions regarding when past mutations must have occurred and from what clades, etc.
Originally, there was only a single haplogroup, but over many thousands of years, others have mutated as populations migrated and became variously isolated or interacted.
That’s a hugely simplified overview, one I’m probably barely qualified to give. But I recommend the Raff and Bolnick paper above along with books by Reich and Rutherford. All three are written in very approachable language without talking down to the reader.
“Who We Are and How We Got Here,” David Reich (https://amzn.to/2Lza55U)
“A Brief History of Everyone Who Ever Lived,” Adam Rutherford (https://amzn.to/2O8j4wK)
You can find the Raff and Bolnick paper by just highlighting the title above and pasting it in a Google search bar.
So without the many bacteria in our digestive system we would not survive. Would you say that their DNA is also human?
Very few people understand the difference between mtDNA and nDNA. That’s why I took so many pages to try to describe the difference. It is necessary to distinguish mtDNA research from nDNA. Back in 2000 I coauthored a text on obesity with an MD specializing in obesity. In it, we went into great detail on mtDNA since it is central to a lot of the best diabetes research. I am very aware of cherrypicking and confirmation bias and selective perception…. it happens on both sides. Denisovan Origins is a theoretical scenario. That was made clear several times.
In essence you really do the same thing you accuse me of when you dismiss the Brazil DNA with the simple words: “The aDNA signals of Austral-Asian ancestry in Brazil are more likely the result of the same land bridge route.” Here’s the point. There is no proof of that. What is left then is what was likely and one’s preferred hypothetical explanation. You prefer Beringia. OK. That’s fine. I go with the S. American archaeologists on it. My belief–and I do mean belief–is that S. American archaeology findings have not been given a fair analysis here. In essence, after the very long Clovis-First fiasco, I and my colleagues–and many, many others–no longer trust mainstream American archaeology. That first assertion you made about us relating that the Solutreans taught the “Indians” mound building–which we never wrote–was a confirmation of why we distrust. That might have just been a misreading or maybe you missed that. But attributing it to us was inaccurate, for whatever the cause. If I made errors in the mtDNA sections of the book, they are certainly mine. I tried to keep it simple for the intended audience and I even wrote that it was probably “too much” for the average reader. But I have yet to find any errors in the DNA sections. If I do I’ll certainly acknowledge them. I’ll acknowledge anything I got wrong once I become aware of it.
The Solutrean Hypothesis was not begun by us. It was begin by archaeologists. It is in the book because it makes sense in explaining the data.Yes, I “sort of” wave my hand and say that in the thousands of DNA studies that go back to the 1980s, any scenario can be supported. Only because that is true.
I have issued about 8 books on mounds and have written again and again, that the ancestors of the Native American tribes built them all. However, I see the ancestors of Native Americans came from a lot of places and over a vast time period. Go to museums in Central and South America and see what their accepted facts are about the First Americans. And the “giants” encountered in S. America–who definitely were not Solutreans. For the readers of such books it boils down to who to trust. I understand and easily accept that many of you don’t trust us, but we knew no matter what we related, you would never trust us.
Thanks Dr. Little for the reply to Carl’s assertions that you, 1.) cite racists sources in your books, 2.) are a racist, or 3.) try to cover up for your racism or that of others. You won’t get too much out of Carl in the way of an apology or admission of error, but good ole Carl will sidestep your questions or even delete your posts “if they don’t feel right”. Again, thanks for defending yourself against the rampant tribalism and double standards that exist here…
Unlike yourself, I suspect Little isn’t one to invent pejorative versions of the names of others who comment here. It doesn’t seem to be his style. And those are the only posts (other than outright spam) that I delete.
However, having gone back and re-read some of Little’s section of the book, I probably should edit my post a little. While I came away with the distinct impression that he was saying Solutrean elite-giants were teaching plain brown natives how to build mounds, it is true that he acknowledged mound-building in the Americas prior to the his believed incursion by Solutreans. I’ll make the edit when I have the time. If I don’t forget. I still see the overall theme as a racist conclusion in many implications. And one that white nationalist and certain white religious elements will favor.
Carl Feagans states: “…Are Collins and Little racist? Probably not intentionally. But not all racists are card-carrying members of the KKK or NAZI’s or any number of other extreme notions you can think of…”.
But even if they’re not “intentional racists”, they’re still suspect, right Carl? Even if you can’t quite pin them down, or find their KKK cards, these fellows are just a little suspicious, don’t ya think? I don’t think you’re a Nazi, Carl, but any real Nazi would be proud of your tactics here. You slander and libel people, and it’s no big deal to you….
Carl states: “…Unlike yourself, I suspect Little isn’t one to invent pejorative versions of the names of others who comment here. It doesn’t seem to be his style. And those are the only posts (other than outright spam) that I delete…”.
Compared to what you’ve called others here, I think my posts are actually pretty lame, unless of course you count that one time use of the word “turd” that caused you morality sensor to go into hyper drive….
On the “trust issue”, and the battle between alleged mainstream and alternative archaeological views: this is a valid point and the main reason for it is because the former cannot ever admit their mistakes, like the Clovis First/Only Model, which Hancock takes to task in his latest book, America Before. It’s clearly wrong, we now know, and no amount of backtracking will ever change that. If that so widely held notion is/was wrong, after being so rigidly defended, what is else is wrong?
The problem with your assumption is that there is no “mainstream” and “alternative” when it comes to archaeology. One is either doing scientific archaeology or one is not.
Pseudoscience/pseudoarchaeological proponents like Hancock, Collins, and Little will always pretend they’re an “alternative.” But the reality is that if they’re approaching archaeology scientifically, then they are the same as their invented “mainstream.”
The Clovis first model was just that: a model. An hypothesis that was the current conclusion of its time. When data finally reached a point that could be replicated in multiple localities by several lines of evidence that pointed to a new conclusion, the model was revised. I should think it will again be revised and refined over time. In fact, I–along with every archaeologist I know–am banking on it. Otherwise why go into archaeology? If I thought there were no new data to discover, uncover, and revise, there would be no point.
Hancock takes Clovis first to task in his book the way a boxer throws punches at his shadow. As a cult leader, Hancock’s shadow boxing is seen as a tournament of skill, one his followers see him as winning. The reality is that he’s in the ring with himself.
Carl states: “…And one that white nationalist and certain white religious elements will favor…”.
Who specifically, or what “religious elements”, are you referring to? I wasn’t aware these fringe groups, as small as they are, made specific use of archaeological data in their tirades…
The LDS church has gone to great lengths to “prove” how the supposed “Lost Tribe of Israel” arrived in the New World and even built the mounds. This notion is today popularized by publications like Ancient American magazine and the anthologies of articles from this magazine published every year or two by former American Nazi Party leader Frank Joseph (believe it or not, he’s actually the guy the writers of the Blues Brothers based the head Nazi–played by the late Henry Gibson–on). And the notion for Mormons isn’t new. The Newark Holy stones fraud was perpetrated in the 1860s to establish a Jewish presence in Ohio associated with the Newark mounds.
The Solutrean hypothesis itself isn’t a racist hypothesis. It’s a genuine scientific hypothesis. The problem is that the data are lacking when it comes to finding support for it. Relying heavily on this hypothesis create a narrative to explain some aspect of Native American (North or South) culture is irrational, without merit, and plays directly into the hands of white nationalists.
There is a 2010 novel dedicated to “the real Native Americans” that portrays Solutreans as the first inhabitants of North America who were killed off by savage invaders from Beringia. The novel’s author is a well-known white supremacist. Stanford and Bradley spoke out against white supremacist claims about their work, as well they should. This is why I’ve been very critical of Collins’ and Little’s irresponsible use of aDNA in their book. They appear to carefully craft discussion about haplogroup X and its sub-clades in a way that equivocates anything that is “X.”
This is dishonest at worst; ignorant at best. Well, at worst, it’s racist if we assume the intent is to inject white people (or Europeans) into the pre-Columbian peopling of the Americas. But I don’t really think that’s their (Little and Collins) goal. So I’ll settle with I truly don’t know if they’re ignorant or dishonest. But modern haplogroup X2 in Orkney has nothing to do with ancient haplogroup X2a in the Americas. While there isn’t yet evidence to show *which* route ancestors of X2a took to arrive in North America, there is better reason to suspect it was via Beringia than an Atlantic route. And this is due to archaeological sites present and aDNA indicators from other haplogroups via Beringia and none via the Atlantic.
I, along with every other archaeologist I know, will cheerfully accept evidence of an Atlantic route. But until then, there is no good reason to think it’s true.
Carl: Perhaps you can correct me, but is the LDS church considered by anyone other than you to be a hive of White Nationalists? I don’t think they are, and I’ll bet if you check the Southern Poverty Law Center’s site on hate groups, the Mormons aren’t listed; so that’s a bad candidate in which to prove your point. Like most who dabble in things archaeological, I’ve heard of the Newark Holy Stones, and a whole collection of similar finds, most now considered frauds, but again, I know of no connection between them and any modern “hate” group. So again, a poor choice. If you look at the stats, even from a left wing site, you’ll find the American Nazi Party to be a very small, to say the least, group. They exist, for sure, like the American Communist Party, but both are nearly extinct. I realize you want to paint any archaeological opinion other than your own in the darkest terms possible, but I think you’re over reacting.
Surely, you don’t expect someone to give you a complete and unabridged accounting of white supremacy, past and present, of the United States in the comments section of my meager blog? I recounted a couple quick examples from the top of my head. You can minimize racism and white supremacy in the U.S. all you like. I’ll not get in your way. And I hope you’re right. I hope it is, indeed, a small aspect of modern culture.
Carl states: “..The problem with your assumption is that there is no “mainstream” and “alternative” when it comes to archaeology…”.
There are alternative opinions, wouldn’t you agree? Do all archaeologists always agree on everything? If they did, I don’t think they’re doing science anymore; it’s more like a cult or a religion where no outside opinion can be tolerated.
Of course archaeologists have differing opinions. But they all base them in science. Sometimes these different opinions compliment each other. Sometimes they don’t. But they’re all rooted in scientific principles and have varied degrees of evidence. In short, they are all archaeology. Not mainstream. Not alternative. It would be like the utterly stupid notion that there might exist some “alternative medicine.” Either something is medicine or it isn’t.
Carl states: “…The Clovis first model was just that: a model. An hypothesis that was the current conclusion of its time…”.
Who first proposed the Clovis First Model, and when? My guess is a mainstream archaeologist and not a fringe (in your view) author. Am I correct? Secondly, is the Clovis First Model now known to be correct, or is it false? Simple questions I think….
Precisely *who* is irrelevant. I’m sure it was probably an archaeologist. Clovis first is absolutely good science in the same way that the Solutrean hypothesis is absolutely good science. That it is now seen as wrong in no way invalidates it as a scientific hypothesis. All science is, necessarily, provisional. I really don’t understand what your hangup with Clovis is. It wasn’t even the accepted model by the time I officially entered the field of archaeology. In fact, that it was an hypothesis overturned is something I celebrate as an archaeologist. I absolutely love being part of a profession willing to adapt and change with evidence.
Carl states: “…Hancock takes Clovis first to task in his book the way a boxer throws punches at his shadow. As a cult leader, Hancock’s shadow boxing is seen as a tournament of skill, one his followers see him as winning. The reality is that he’s in the ring with himself…”.
What percentage of mainstream archaeologists would you say once supported the Clovis First Model? 10%? 90%? What present percentage now support it? Hancock’s main point is this; if you were a credentialed archaeologist who supported the Clovis First Model, you were simply wrong.
Here is Hancock’s original, word for word statement, taken from page 27 of his book, America Before:
“…At the outset of the 20th century many scholars took the view that the America’s had been devoid of any human presence until less than 4000 years ago…”
In your opinion, is that statement true or false?
LOL. Why would I give a shit about the conclusions of scholars at the outset of the 20th century?
At the outset of the 20th century many scholars took the view that contamination of the “Nordic race” was a threat to be warned of, and Ludwig Woltmann creates a journal called Political Anthropology Revue” to perpetuate this warning.
At the outset of the 20th century many scholars took the view radium water and mercury could heal blindness and syphilis.
At the outset of the 20th century many scholars took the view that there was an “ether” acted as a transmission medium for light. An idea Einstein demolished in 1905.
Undoubtedly, there will be things we all believe scientifically true that will be looked upon as quaint in a few hundred years. That science is provisional is a good thing.
Carl states: “…Not mainstream. Not alternative. It would be like the utterly stupid notion that there might exist some “alternative medicine.” Either something is medicine or it isn’t…”.
You have a tendency to express things in an “either/or” fashion; I’m sure you took history courses in college and you’re acutely aware of such statements of “either your a loyal American or you’re not”, or more recently as George Bush stated, “your either for us or against us”. Do you really believe it’s all just that simple? I recall, years ago watching a small debate between 2 credentialed doctors on the subject of breast cancer and the best way to proceed. One was an advocate of the radical mastectomy and the other a lumpectomy approach. Even though they were in vehement disagreement, don’t you think both believed science was on their side, and that surely both arrived at their conclusions based on science? I’m only trying to argue that the “either/or” approach is severely flawed….
Carl: “…Surely, you don’t expect someone to give you a complete and unabridged accounting of white supremacy, past and present, of the United States in the comments section of my meager blog? I recounted a couple quick examples from the top of my head. You can minimize racism and white supremacy in the U.S. all you like. I’ll not get in your way. And I hope you’re right. I hope it is, indeed, a small aspect of modern culture…”.
Carl, I majored in history (emphasis in US history), BS degree from Charleston Southern University, 2000, so I think I’ve got you beat there, knowledge wise. Do you think I am trying to minimize the history of racism in the US? Are you shortly going to accuse me of being a racist? I was merely responding to what you perceive to be groups associated with White Nationalism, in which you included the LDS church. And yes, the US Nazis/Communists/Klan are almost extinct…
Carl states: “…You can minimize racism and white supremacy in the U.S. all you like. I’ll not get in your way. And I hope you’re right. I hope it is, indeed, a small aspect of modern culture…”.
Or is it you, for purely political reasons, over-emphasizing racism is the US? I hope you aren’t doing that. Or were you perhaps raised in a back woods, racist, Kentucky home, and you are now “projecting” just a bit? I hope not…
Assumptions, assumptions… 🙂 In 53 years, I’ve lived in 4 countries and 8 states and I’m on my fourth and final career. I detest politics and I have a lot of faith in people. But I’ve seen my share of racism and bigotry from the tiniest degree to the most extreme.
Carl states: “…Precisely *who* is irrelevant. I’m sure it was probably an archaeologist…”
Of course the “who” is irrelevant, no one said it was. And yes, almost for certain a mainstream archaeologist, not a fringe author. That’s all I’m getting at.
Carl states: “I really don’t understand what your hangup with Clovis is. It wasn’t even the accepted model by the time I officially entered the field of archaeology. In fact, that it was an hypothesis overturned is something I celebrate as an archaeologist. I absolutely love being part of a profession willing to adapt and change with evidence…”.
I don’t have a “hang up” with it; apparently you do. The Clovis First Model is now known to be 100% wrong, false, bogus, fraudulent. That’s all Hancock is saying. Is that embarrassing to you for some reason?
Hancock’s obsession (and yours, apparently. You keep bringing it up, not me) is hardly embarrassing to me. Though it should be to him.
Clovis first is wrong. It is false. But it isn’t bogus nor was it fraudulent (these are terms that imply an intent to deceive). It was good science. Until it wasn’t. That’s all. Science is provisional. New data showed that new conclusions needed to be accepted. Now they are. For decades.
“…Undoubtedly, there will be things we all believe scientifically true that will be looked upon as quaint in a few hundred years. That science is provisional is a good thing…”
Shit, Carl, we actually agree on something!
I bet we agree on far, far more than you think.
Carl states: “…Assumptions, assumptions… ? In 53 years, I’ve lived in 4 countries and 8 states and I’m on my fourth and final career. I detest politics and I have a lot of faith in people. But I’ve seen my share of racism and bigotry from the tiniest degree to the most extreme…”.
Nope, not assuming anything. I’m 56, raised in SC, in my last career position also, was in the US navy for 9 years and have lived in Tenn, Mississippi and California during that span; yes, have witnessed racism but never anything “extreme”….but, that’s just my life history, not saying it is the norm. But I have witnessed great cooperation and acts of compassion between “races”…
Carl states: “…I bet we agree on far, far more than you think…”.
Certainly, no doubts about that. I read your blog even if I don’t agree with all of it. I read Hancock’s books and certainly don’t agree with all of what he says either. I just wish sometimes that the disagreements between the two “sides” wasn’t so bitter sometimes….:)…
Hey Carl: It’s probably a normal thing, since the two subject areas are so obviously intertwined, but a lot of history majors have a great interest in archaeology as well, hence some of my interest in your blog. Sadly perhaps, or maybe irony is more correct, I think my introduction to the latter subject was running across a copy of Chariots of the Gods while in my early teens. Of course, now I know a large portion of what von Daniken wrote was false, but anyway, it did get me interested….:)
It is funny how you pull out the SJW, “muh racist” card, because someone said, a group may have taught another group. Did you pull out the racist card, when Andrew Collins said, the Denisovans may have taught the WHITE people of Anatolia, South eastern Europe and Iran, how to do certain things.
You people are are truly pathetic. Looking for every chance to virtual signal what great people you are and how much White guilt you have.
Talk about the king of cherry picks. I think they should sue you for libel.
I know they get lots of things wrong and some of the stuff is out there, but you are disgusting.
I don’t really know what some of what you wrote means. I’m unfamiliar with the abbreviation “SJW” and only assume the first letter is “Solutrean,” so… “Solutrean Journey West?” This isn’t a familiar term to me. Nor is “muh,” so I really don’t know what you’re saying. It does seem, however, that you didn’t read what I wrote and that you’re having a knee-jerk reaction to a few keywords you picked out.
More importantly, I think you’re probably not familiar with what racism really is and assume it’s simply an accusation that people like to fling about to look cool. Not all racism is made of overt actions or words intended to oppress a given ethnic class. Many times, ideas and notions themselves can be racist without intent. Is either Collins or Little a racist? I really don’t know. I like to think neither is. My critique is of their work and the ideas they present. These, I believe, are racist for the reasons I’ve outlined.
Regardless, thank you very much for the time you took to provide feedback in the comments section. Clearly you’ve disagreed with my conclusions in regard to this book (I’m assuming you read it), but hopefully you found one or more of my other posts more enjoyable or informative.
And here we go again….Argumentum ad Clovisum
My new concept for describing the fringe’s tendency to insert Clovis First, albeit incorrectly, into just about any argument that they have with someone who doesn’t buy into Hancockian nonsense or that of other fringers.
But at least no lectures on the omniscience of electrical engineers, the persecution of Gallileo, or the infallibility of Schoch’s “research” so things could have been worse.
Carl, I will say it yet again. You are a saint.
Well, James, those mean old electrical engineers at least keep your lights burning at night, otherwise how would you ever find your two coloring books and and your “research papers” from a career as an unlicensed pharmacist?
If you reduce Hancock’s main argument down to it’s absolute barest minimum, what you will find is a single, simple, proposition; that mainstream archaeology is not only fallible, but in some cases utterly wrong It’s so plainly a demonstrable fact, that only someone who views mainstream archaeology as a type of religion cannot readily admit it. That the Clovis First model was deeply flawed and now discarded is an excellent example of that. But, as in many human institutions, I can understand no one likes having their authority questioned; after all, if the Clovis First Model was vehemently defended for *decades*, what else in current archaeology is completely wrong?
No archaeologist I know disputes that. In fact, we professionals in the field see these as attributes. That we admit fallibility and the willingness to revise with good evidence is the strength of science. In fact, as a professional archaeologist, it is my hope that we revise many more assumptions over time with good and better evidence. Unfortunately for Hancock cult-followers, these revisions will likely be as mundane as the Clovis stuff. Pushing back human migration from 13 ka to about 24 ka, is exciting to professionals, but really doesn’t change a lot about what we knew from that 13,000 years to now. It does, however, begin to add some additional chapters. The lunacy Hancock and his cult-followers suggest is inconsistent and irrational… a.k.a pseudoarchaeological and pseudoscientific.
LOL. “High technology” and “Atlantis in America” and all that other hocus pocus of a “lost” civilization that left no trace.
Carl Feagins:
Just a heads up. Charleston Southern University was formerly known as Baptist College of Charleston a small private institution associated with the Southern Baptist Convention. Not the best place to get quality undergraduate instruction in the history of race and racism. It certainly doesn’t provide a high intellectual ground from which to argue with someone with an advanced degree in anthropology. It helps to explain why gleaner is overly sensitive to any discussion of the various forms that racism can take besides people burning crosses. These institutions are not exactly bastions of anthropological teaching and research for obvious reasons, unless one is interested in things like “biblical archaeology.” CSU does not appear to offer a single undergraduate course in anthropology. It helps to explain Gleaner’s ignorance and mistrust of anthropology and often outright hostility toward it. It looks like all of gleaner’s criticism of anthropology are drawn directly from talking points from people like Graham Hancock instead of his own reading and understanding of anthropological writings. Also helps to explain why gleaner is willing to aggressively defend research that would appear to lend credence to the belief in the existence of giants in ancient history. Most of these types of schools had to be dragged kicking and screaming into the 20th century. They are still working on the whole 21st century concept. I attended a small conservative Lutheran College after a couple years of imprisonment at a SBC affiliated academy. The Lutheran school which was pretty far ahead of the game with one fulltime anthropology faculty and two adjunct lecturers. Compared to a southern bible college it still had its share of people who don’t know what they don’t know and are very happy with it. So take that for what its worth when dealing with someone whose intellectual insights come from attending a southern bible college two decades ago when many people in South Carolina still couldn’t figure out who had won the civil war and thought that calling a black person Mister demonstrated that they were colorblind.
Correct me if I am wrong, but my understanding is that the Clovis First theory held that small groups of hunters were present in the western hemisphere around 13,000 years ago. Subsequent research has simply demonstrated that small groups of hunters were present in the western hemisphere a few thousand years earlier. Certainly an interesting development but I fail to see this how this can be spun as something other than a new finding based on further research and further refinement of dating technology. Am I missing something here? Needless to say my question is for Carl Feagins.
Happily Lapsed Baptist:..so you can’t engage with my arguments, so instead you attack the school I went to, an entire region and state, and then me; wow, talk about a complete misunderstanding of science and history and a hypocrite to boot. But, you probably know so little about US history there’s really no other recourse.
Sure, Carl, you see I’m not making any fantastic claims here, and I’ve said absolutely *nothing* about “Atlantis in America” or “High Technology”, so I don’t really know where’s that coming from. Perhaps, if you would give people a little more credit for being able to think for themselves, you might find that I not only don’t adhere to everything Hancock says, I also hold you to the same standard; I don’t accept everything you write here as the absolute truth either, which I think you would agree is a good thing, correct?
You mentioned Hancock. This was in reference to him.
Happily Lapsed Baptist states: “…Correct me if I am wrong, but my understanding is that the Clovis First theory held that small groups of hunters were present in the western hemisphere around 13,000 years ago. Subsequent research has simply demonstrated that small groups of hunters were present in the western hemisphere a few thousand years earlier. Certainly an interesting development but I fail to see this how this can be spun as something other than a new finding based on further research and further refinement of dating technology. Am I missing something here?..”.
Yes indeed, you are missing something here, although I don’t know if it’s a lack of education on your part in any relevant field, or you simply didn’t read prior posts in this thread. So anyway, I’ll use some of my valuable time to “fill you in”; now try to keep up!
Graham Hancock states in his book “America Before”: “…At the outset of the 20th century many scholars took the view that the America’s had been devoid of any human presence until less than 4000 years ago…”. Do you comprehend the importance of that statement? That means at one time the consensus was homo sapiens sapiens had only inhabited the America’s no earlier than 2000 BCE. The Clovis FIrst Model then pushed that date back farther, prior to 10,000 BCE. That’s an error of 8000 years. Now with new evidence, that date might be pushed back even further, as Carl has stated “to 24[000] ka”. That’s another error of 10,000 years. Somewhere in the archeological establishment someone has made some egregious mistakes. To go from 2000 BCE to possibly 24000 BCE, or even further is substantial. What is strange though, is that apparently, judging by your prior posts, you weren’t aware of any of this…
ka means “kilo annum” or a “thousand years.”
And the incorrect dates science held for the peopling of the Americas wasn’t exactly an error. It was a provisional conclusion. One that changed with the advent of better data. One that will likely change again with newer data. Some of this data were simply not available when the initial conclusions were provisionally made. It was advancements in physics and genetics that have allowed for better data collection, testing, and replication along with better methods in archaeology based on improved theoretical frameworks.
Hancock’s real beef is that he isn’t taken seriously by professional archaeologists and other scientists. And its this professionalization of archaeology, a process that began in the early 20th century, that has excluded pseudoscientific thinkers like Donnelly, Velikovsky, Sitchin, Hancock, Collins, et al from the table. This causes resentment and the invention of terms like “mainstream” to describe what is actually real, scientific archaeology and the attempt to create an “alternative” for that archaeological thought which is both non-professional and largely non-scientific. I say largely, because there is often a veneer of science. A “cargo-cult” version of archaeology, if you will, that *pretends* to be scientific by literally donning the lab coats and adopting methods until they reach a point that the methods don’t produce the data they want.
Happily Lapsed Baptist states: “…Just a heads up. Charleston Southern University was formerly known as Baptist College of Charleston a small private institution associated with the Southern Baptist Convention. Not the best place to get quality undergraduate instruction in the history of race and racism…”.
I also attended West Hills Jr. College and Chapman University in California. But I’m curious, what university would you recommend someone attend for “quality instruction” in race and racism? This out to be a real howler…
Carl states: “…You mentioned Hancock. This was in reference to him…”. I understood the reference, what I don’t understand is the painting with such a broad brush, that anyone who makes even a passing mention of Hancock must in some way be in 100% agreement with him, even to the point of belonging to a “cult”. So far, I’ve seen no conclusive proof from Hancock or anyone else that points to an unknown “high culture” that is now lost to history…
I agree that Hancock is not taken seriously by most archeologists or scientists and I’m sure he knows that; he interviewed Dr. Goodyear about the Topper Site and others of similar stature and I don’t think it hurts anything to at least ask the questions. And as far as I can tell, none of those archaeologists were hostile to him…
Well Gleaner63 feel free to tell the electrical engineers that I said thanks for the hot showers, cold cream soda, and the use of computers that made it so much easier for me to publish peer reviewed journal articles on substance use and abuse and print out the syllabus for my Drug and Alcohol Studies college course. Of course that doesn’t make me less skeptical of claims that electrical engineers have long had all the answers when it come to the nature and properties of electricity. To expand on what others have previously noted:
https://www.quantamagazine.org/newfound-tetraquark-fuels-quantum-debate-20140827/
https://www.nytimes.com/1979/02/13/archives/new-quarks-stir-debate-on-basic-laws-of-nature-fundamental-building.html
Now here is where you will try to quibble over semantics to save face, but I consider the matter closed. Thanks Mr. Quark and Mr. Preon.
While we are on the topic of thanks and race maybe a shoutout to anthropologist who have spent the last century working to undercut the “scientific” support for racism and white supremacy. How about the archaeologists and forensic anthropologists who help to investigate war crimes and other crimes and work to recover and identify our war dead. I would suggest that you go back and take a long look at the pioneering work of people like Boas and Herskovits and work your way forward but you and I know that just ain’t gonna happen. Ditto for the doctors who you undoubtedly have significant faith in despite the fact that they were once WRONG about things like bleeding people and not washing their hands between delivering babies, and they still have to carry a lot of malpractice insurance. Those frauds!!
Graham Hancock has an “argument” about Clovis? I though that he was just a journalist asking questions LOL. It takes a lot to impress me, but your utter lack of insight or understanding as it relates to this topic and its relationship to how science works is truly impressive. I could recommend some published papers on pre-Clovis research going back almost 40 years. You know back when Hancock says that dogma and conspiracy prevented anyone from talking about it. But you and I know ya just ain’t interested in anything that doesn’t parrot a Hancock talking point.
Speaking of pharmacists I gotta make a Sudafed run. Or maybe that fraud of a pharmacist was running a major con job on me when he suggested that I give it a try for this head cold. You know those bastards have been known to be wrong so why trust any of them now? Maybe I should instead visit one of Hancock’s shaman buddies and get a little something-something instead. May not help the congestion but I will be too busy communing with spirits that may or may not be my imagination telling me how the pyramids were built to notice it.
Buenos Nachos amigo.
James Ford states: “…Of course that doesn’t make me less skeptical of claims that electrical engineers have long had all the answers when it come to the nature and properties of electricity…”.
I’m not sure what your hang up or phobia is about electrical engineers but it must be pretty severe; maybe you should write a paper on it and submit it to IEEE and see if they will publish it? But look, I can understand that “drug researchers” and electrical engineers (I believe you actually confused an EE with an “electrician” a few posts back, LOL) might speak different sorts of language; so let me straighten that out for you. EEs have to deal in regularities, even if, ultimately, the “nature” of those regularities is not completely understood. For example, that 110 volts AC powering your lights while you are attempting to light you bong is always going to be 110 volts (plus or minus a few volts) or your lights will soon be off; it’s not going to suddenly switch to DC, or 220V or 440V. Meanwhile, over in the Clovis First department, the stunning revelation that an error of 20,000 years may have been discovered is causing a lot of concern. If I made such an error on my job I’d be gone tomorrow. But look at the bright side, you can always claim that anything you say is “provisional”, which is just another way of saying “we don’t really know what the hell we’re talking about”
James Ford states: “…Graham Hancock has an “argument” about Clovis?…”.
As is now admitted by Carl and the rest of the “mainstream” archaeological community, the Clovis First Model has now been regulated to the category of pseudo-archaeology, the fringe, and ancient astronaut theorists. It was wrong by tens of thousands of years, despite a vigorous defense by what are now known to be “cranks”. It’s funny you know, how yesterday’s “science” is today’s “pseudo-science”….
Either you’re complete uneducated to the point of not capable of basic reading and comprehension or you’re intentionally misconstruing everything I’ve said. Please quote where I say what you write above.
James Ford states: “…It takes a lot to impress me…”.
Yeah, I can see that. Your first homemade bong and the paper you wrote about it, did you at that point consider yourself an inventor, mechanical engineer and peer review expert? Just funnin with you..:)….
Happily Lapsed Baptist: How odd, that in your post you assail an entire institution (Charleston Southern), a religion (Christianity), a state (SC), and a region (the American South), and after all that, you then lecture us about the evils of racism. Are you really THAT blind to your own prejudices or is that your own views directed towards millions of others is a perfectly acceptable type of prejudice? In other words, you don’t really, ultimately, have a problem with racism, prejudice or stereotyping at all, just so long as it’s directed at the right type of people (in this case, white, Christian, Southern, conservatives)……
Carl states: “…Either you’re complete uneducated to the point of not capable of basic reading and comprehension or you’re intentionally misconstruing everything I’ve said. Please quote where I say what you write above….”.
Carl states: “…Clovis first is wrong. It is false. But it isn’t bogus nor was it fraudulent (these are terms that imply an intent to deceive). It was good science. Until it wasn’t. That’s all. Science is provisional. New data showed that new conclusions needed to be accepted. Now they are. For decades…”.
Not misconstruing what you said, merely making this point: when Hancock makes a mistake, he’s immediately accused of being a pseudo-scientist or a pseudo-archaeologist, or something far worse. Basically, he’s given no margin of error; maybe he simply didn’t understand the data, maybe he read something out of context, maybe, just maybe, being human, he made a simple mistake and there’s nothing more nefarious behind it. So now that the Clovis First Model has been shown to be unbelievably wrong, by a wide margin, why can’t I apply the same terms that you use for others? In short, when a mainstream scientist makes a mistake (like Carl Sagan), what ensues is always a case of special pleading or a double standard…
And therein lies your ignorance. Willful at this point since it’s been explained to you in many comments over several posts. Clovis first, as a hypothesis was certainly not “unbelievably wrong” if you’re educated in science and the methods of science. It was the best explanation possible for its time until new evidence came along to show it wrong. That you continue to beat that long dead horse shows your willful ignorance. I see little reason to continue a conversation or dialog with someone unwilling to overcome this ignorance. Cheers.
Here’s a great example of a double standard, from wikipedia:
“…In Intelligent Life in the Universe (1966) astrophysicists Iosif Shklovsky [Shklovskii] and Carl Sagan devote a chapter to the argument that scientists and historians should seriously consider the possibility that extraterrestrial contact occurred during recorded history; however, Shklovskii and Sagan stressed that these ideas were speculative and unproven.[20] Shklovskii and Sagan argued that sub-lightspeed interstellar travel by extraterrestrial life was a certainty when considering technologies that were established or feasible in the late 1960s;[21] that repeated instances of extraterrestrial visitation to Earth were plausible;[22] and that pre-scientific narratives can offer a potentially reliable means of describing contact with aliens…”
If true, how is it that Carl Sagan is never taken to task over these positions, but Von Daniken is raked over the coals..? Was Sagan, suddenly a pseudo-scientist, at least for a day, before he came to his senses? Can one in fact be a scientist on Monday, a pseudo-scientist on Saturday night, and then suddenly return to having “correct thoughts” come Sunday morning? Or is one a pseudo-scientist for life, once the label has been applied?
Jame Ford referred me to this article: “https://www.nytimes.com/1979/02/13/archives/new-quarks-stir-debate-on-basic-laws-of-nature-fundamental-building.html”
Since, by your own admission, you are vastly more educated than me, could you explain how this article contains anything that changes how an EE does their job? Just curious…
Gleaner:
You are the one who specifically cited a degree from Charleston Southern University as the basis for speaking authoritatively here. So that’s what’s on the table. Now that some context has been provided for your undergraduate education there you want to shift the conversation to other schools. One is yet another small private religious school and the other is a two-year community college that one can presume only offers basic introductory courses. Not exactly the stuff that stands out on a resume. If you had any sort of significant in-depth coursework in the relevant areas at these schools I assume that we would have heard about it by now. As for institutions that offer coursework in matters of race and racism, just about any medium to large public or private school has departments, programs, and faculty that offer significant coursework that provides focus on these topics: for example, anthropology, sociology, ethnic studies, African-American Studies, Pan-African Studies, History, Race and Ethnic Studies. Well represented in nearly all if not all Big Ten, SEC, and ACC programs just to name a relative handful. Plenty of small liberal arts colleges that fit the bill too, including some Church affiliates. Millsaps which one of my professors attended comes to mind as a fine school with a diverse relevant curriculum. But then again the Methodists aren’t wrapped nearly as tight as the southern Baptists when it comes to things like this. Got a buddy who thinks highly of what they do at Rhodes College in terms of a legitimate liberal arts education and the Presbyterians are fairly low on the scale in terms of being tightly wrapped.. As it relates to South Carolina, The University of South Carolina comes to mind, as does Clemson where one of my former professors is on faculty, as does the College of Charleston where one of my cousins attended. As for specific quality of programs, various professional organizations as well as publications such as the Chronicles of Higher Education provide annual rankings of various schools, departments, and programs. Even institutions that may not be well ranked may have individual faculty scattered across programs and departments that are highly regarded. Pays to do your homework when picking a school. That’s the long answer that you undoubtedly find to be a howler. Now for the real howler. The short answer is that a monkey could draw a name from a bucket and pick a better relevant institution than Charleston Southern. Chapman sounds like a step up from CSU and the people running it aren’t exactly snake handlers. But based on your comments here it doesn’t appear that you took advantage of any relevant learning opportunities that may exist there. That’s clearly why you don’t even have a proper educational frame of reference for grasping most of what is being discussed here but will argue endlessly anyway.
Carl, it’s your blog, so you can do whatever you like, you can ban me, delete my posts or refuse to talk with me, that’s up to you. I majored in US History and minored in Secondary Education; although the majority of my courses were in those two areas, I also took Astronomy, Geology, Botany, Anthropology and Biology. Despite another of your readers degrading the college I graduated from (Charleston Southern), I took my Intro to Anthropology at The College of Charleston, which is not known to be affiliated with creationists in any way. So while I don’t have your complete background in science, I do work in a technical field and you’re completely off base to call me “willfully ignorant”. But hey, if it makes you feel any better, and think it someone makes your arguments stronger, then have at it. I didn’t come here for you to “explain” anything to me; after all, having never heard of you, and I believe you’ve never written a book on archaeology, do you honestly think I believe you’re some kind of world known expert on this subject? I don’t. On the Clovis First Model, which you readily admit has been canned, I already knew that, so again, you didn’t reveal any new knowledge to me. I hate to quibble with you over phrases like “unbelievably wrong”, but in the dates department, it certainly seems that “Clovis First” was wrong by a wide margin, and could get worse. I understand you’re defending your profession, that’s a normal thing to do. I don’t really have a dog in this hunt in that regard. Anyway, don’t take it so personal, people are going to disagree and that’s a good thing, I think.
Archaeological sites that antedate Clovis that are well documented include:
Bluefish Caves, Yukon, Canada (24,000 yr BP)[54]
Pedra Furada, Piauí, Brazil (10,500–12,000 yr BP; possibly >50,000 yr BP but this is disputed)[55][56]
Topper, South Carolina, US (16,000–20,000 yr BP; possibly 50,000 yr BP but this is disputed)[57][58][59]
Meadowcroft Rockshelter, Pennsylvania, US (16,000 yr BP)[60]
Buttermilk Creek Complex, Salado, Texas, US (15,500 14C yr BP)[10][61][62]
Cactus Hill, Virginia, US (15,070 14C yr BP)[63]
Monte Verde, Chile (18,500 to 14,800[64] 14C yr BP)[65][66]
Saltville (archaeological site), Virginia, US (14,510 14C yr BP)[67]
Taima-Taima, Venezuela (14,000 yr BP)[68]
Manis Mastodon Site, Sequim, Washington, US (13,800 yr BP)[69]
Connley Caves, Oregon, US (13,000 yr BP)[70]
Page-Ladson, Florida, US (14,550 cal yr BP])[71]
Lapa do Boquete, Brazil (12,070 ±170 14C yr BP)[65][72]
Paisley Caves, Oregon, US (14,300 cal yr BP)[73]
Tanana Valley, Alaska, US (13,000–14,000 cal yr BP)[74]
El Abra, Colombia (12,460 ±140 14C yr BP)[65]
Nenana Valley, Alaska, US (12,000 yr BP)[75]
Tibitó, Colombia (11,740 ±110 14C yr BP)[65]
Tagua-Tagua, Chile (11,380 ±380 14C yr BP)[76]
I would say this data confirms, or will confirm, that the “Clovis First Consensus” was more than just “wrong”; it was really wrong. And yes, I’m fully aware of how new evidence in the best traditions of the scientific method are needed to overturn older theories; I get that and completely agree with it…
Happily Lapsed Baptist states: “…You are the one who specifically cited a degree from Charleston Southern University as the basis for speaking authoritatively here. So that’s what’s on the table…”.
No I did not; see, you are already lying about something I said, straight out of the gate. I have NEVER made an appeal to my college education as the basis for speaking authoritatively on anything here. I mentioned it because it was a way of showing what I am most familiar with (US history), and what I wasn’t familiar with (anthropology). If you would have read my other posts instead of showing up here with a hammer, you would know that. So you will have to explain you reasons for you unwarranted attacks on me…
Baptist states: “…Not exactly the stuff that stands out on a resume. If you had any sort of significant in-depth coursework in the relevant areas at these schools I assume that we would have heard about it by now…”.
Why? You seem bent on putting me in a box. First you attack me because I went to a small religious school, and now because West Hills Jr. college was a small 2 year college…what’s your point?
Baptist states: “…anthropology, sociology, ethnic studies, African-American Studies, Pan-African Studies, History, Race and Ethnic Studies. Well represented in nearly all if not all Big Ten, SEC, and ACC programs just to name a relative handful. Plenty of small liberal arts colleges that fit the bill too, including some Church affiliates…”
Of the 7 courses/areas of study you mentioned, I took 3 of them; but I never intended to major in ethnic studies, and my main interest was in US history….
Baptist states: “…As it relates to South Carolina, The University of South Carolina comes to mind, as does Clemson where one of my former professors is on faculty, as does the College of Charleston where one of my cousins attended…”
Are you from SC? My dad graduated from Clemson in 1950, and both of my sisters graduated from SC, so I’m quite familiar with both of those schools…
Baptist: “…Now for the real howler. The short answer is that a monkey could draw a name from a bucket and pick a better relevant institution than Charleston Southern. Chapman sounds like a step up from CSU and the people running it aren’t exactly snake handlers. But based on your comments here it doesn’t appear that you took advantage of any relevant learning opportunities that may exist there. That’s clearly why you don’t even have a proper educational frame of reference for grasping most of what is being discussed here but will argue endlessly anyway…”.
Wow, once again, you trash schools you didn’t attend and volunteer info that no one asked for. Assuming you actually went to college (I doubt it), you need to get a refund…LOL…
Baptist states: “…As for institutions that offer coursework in matters of race and racism, just about any medium to large public or private school has departments, programs, and faculty that offer significant coursework that provides focus on these topics: for example..”
A howler for sure; since you don’t get sarcasm, the question was asked purely in jest, and now you go off the deep end of the pool. Even a Monkey could have gotten the sarcasm in my prior post, so here’s some free advice for you; no one, and especially me, gives a crap about your opinions on race or relevant college choices. You pop up here outta nowhere, don’t read the prior posts, misquote or don’t understand what I’m saying, and then you start giving out free advice. You sound a little “thick”. And what was the name again of that mythical college you attended? LOL…
Baptist: I know you won’t “get it”, because your tone marks you as narrow minded and bigoted but not everyone has the same path when pursuing their education. When I was in the military, most of one’s college choices were limited to the general proximity of the base you were assigned to; hence my decision to attend West Hills Jr. college and then Chapman University. When I got out of the navy and moved back to SC, I took courses at USC-Allendale, USC-Aiken, and the College of Charleston, before finally graduating from Charleston Southern. That’s what best fit my situation and career path. And by the way, I’m still waiting on you to tell us where you graduated from..chirp, chirp…
James Ford states: “…While we are on the topic of thanks and race maybe a shoutout to anthropologist who have spent the last century working to undercut the “scientific” support for racism and white supremacy. How about the archaeologists and forensic anthropologists who help to investigate war crimes and other crimes and work to recover and identify our war dead. I would suggest that you go back and take a long look at the pioneering work of people like Boas and Herskovits and work your way forward but you and I know that just ain’t gonna happen. Ditto for the doctors who you undoubtedly have significant faith in despite the fact that they were once WRONG about things like bleeding people and not washing their hands between delivering babies, and they still have to carry a lot of malpractice insurance. Those frauds!!..
Well, interesting to say the least, incoherent at best, but certainly worth a second read if the above was written by someone in an insane asylum. And who knew that most normal folks put their medical doctors in the same category as archaeologists? I can see it all now, in my head, as the doctor is summarizing my latest physical results, and I’m thinking to myself “…but did he ever accept the Clovis First Model as genuine, why or why not…?. I don’t know about you, James, but on the basis of how he answers that question will go a long way toward deciding how good a doc he really is. Don’t misunderstand me; I’m not saying that Cancer Research is unimportant, because surely it is, but it’s not even a close second to the validity of the Topper Site. Welcome back, Mr. Loon….
Gleaner said, “Carl, I majored in history (emphasis in US History), BS degree from Charleston Southern University, so I think I got you beat there knowledgewise.”
A clear appeal to your education at CSU to speak authoritatively. Who is lying here?
gleaner said: “But I’m curious, what university would you recommend someone attend for “quality instruction” in race and racism.”
You solicited information from me and now pretend that you really didn’t because you couldn’t handle the answer. Who is lying here.
If you can’t keep your story straight in these areas then there is really no need to take anything else that you assert here seriously. it’s just a combination of willful ignorance and trolling.
“I’m still waiting on you to tell us where you graduated from.”
I don’t recall being asked for that information although such a question could easily be missed in the many long rambling sermon-like posts you have here. I’ll take a pass on communion and the collection plate now, though. But to answer your question I finished (note FINISHED) my undergraduate career at Southern Illinois University with a B.A. in Anthropology and History, emphasis on archaeology as well as North American and Latin American History. Produced a well-regarded senior history thesis. Earned an M.A. in Anthropology at LSU with advanced coursework and oral and written comprehensive exams in Archaeology, Linguistic Anthropology, Physical Anthropology, and Cultural Anthropology, with cross training in History and Geography. Ph.d. in Historical Anthropology from University of Illinois. Strong concentration in history and ethnography of the U.S. South and race and ethnicity in my graduate work. Published articles and book reviews in Louisiana History, Journal of American Ethnic History, Journal of Cultural Geography, Ethnology, and Southern Journal of Linguistics just to name a few. Also served as a college advisor at two different institutions (one small college, one mid-sized university)who had to have in-depth knowledge of various programs and colleges across the nation to assist in matching students with appropriate quality programs in fields such as anthropology, history, race and ethnic studies, etc. I’m quite confident that I have forgotten more about just about anything discussed here than you ever learned in your entire college career. Unless of course you think that you could keep up with me in a discussion of topics like how the 1870 Federal Population census can be used to critically analyze the concept of hypodescent and racial classification and racism in the US South. Or maybe a discussion of how to develop a semi-structured interview protocol to help develop a database on illicit racism and its influence on observed behaviors in a rural southern community.
However well your education, introductory anthropology course from a bible college and all, may have fit your situation and career path it is not serving you well here. You come across like a 13 year old trying to argue with a table full of adults and thinking that you are really accomplishing something when every one else in the room just sees you as foolish. For example:
I’m no expert on the Clovis issue. But I did learn about it in numerous archaeology classes. Like Carl Feagins and others have discussed here, my memory of how it was discussed by archaeologists does not mesh with Graham Hancock’s portrayal of it. Hancock’s account does not even jibe with what I recall reading about Clovis in high school textbooks. The vast majority of archaeologists had other things on their mind besides whether small numbers of hunters and gatherers with similar technology were in North America 18,000 years ago or 13,000 years ago. Hell, I always thought it would go back much further myself, and still do. But thinking and proving are two different things. The Clovis issue actually involved a very small group of specialists whose thoughts and actions do not necessarily reflect archaeology as a whole beyond illustrating how this stuff is supposed to work. You can’t grasp that and are unwilling to even make the effort. Willful ignorance. New data and refined dating methods caused scientists to rethink the dates of early human occupation and made it a more popular topic. But that is very old news and, again, how science is supposed to work. If you want to continue to suffer under the delusion that this somehow means that archaeologists are frauds or pseudoscientists or even absolutely wrong about Clovis (the model actually involved many aspects beyond just dates and much of this remains correct even after decades of pre-Clovis findings) then perhaps you would be better served seeking other venues to discuss the matter. Simply googling up dates for pre-Clovis sites and ignoring everything else involved in the model just demonstrates your lack of knowledge of the overall subject. You are simply coming across as childish here and no one other than fellow Hancock fans takes you seriousy. Carl has been overindulgent thus far since it is quite clear that you are more interested in trolling or serving as the poster child for willful ignorance than having any sort of actual informed discussion or learning from people who are light years ahead of you in terms of relevant education and actual research experience. Now I have finished my own sermon, albeit a much better one than your’s, and this should keep you raving on your soapbox for the rest of the weekend. That will have to be sufficient for you to continue the self-delusion because I’m not going to waste any more time angering and confusing you with facts and logic. At this point it would just be cruel to continue anyway. Cheers.
Baptist states: “Southern Illinois University with a B.A”
Wow, so you graduated from Southern Illinois? Never heard of them, and certainly nothing to brag about..lol…
On the contrary, SIU has an extremely high reputation among professional archaeologists. I do my fair share of hiring, and when I see SIU on a resume I take note.
Baptist states: “…You solicited information from me and now pretend that you really didn’t because you couldn’t handle the answer. Who is lying here…”.
You are again; clearly my question was in jest, as I said before, but perhaps you are, as I also said, a little “thick”…
Baptist states: “…You come across like a 13 year old trying to argue with a table full of adults..”.
Let’s see; you show up here out of the blue, denigrate colleges you know nothing about, a whole region of people, and to insult their religion you use the term “snake handlers”. But I’m the one who comes across as 13???? And yet, despite all the personal attacks, you still haven’t engaged with a single argument I made….
Baptist states: “…At this point it would just be cruel to continue anyway. Cheers…”.
A cop out, because you’re lying about your education and you can’t engage a real argument, preferring the ad hom route..sure…what an idiot…
Baptist states: “..Gleaner said, “Carl, I majored in history (emphasis in US History), BS degree from Charleston Southern University, so I think I got you beat there knowledgewise.” A clear appeal to your education at CSU to speak authoritatively. Who is lying here?”.
The narrative in these types of threads usually follows a very predictable pattern. If one disagrees with any opinion held by mainstream archaeologists, the usual response is “..well, your just uneducated…”. When you can show that you have a college degree, the charge then changes to “…well, you went to a crappy college…”, or “you don’t understand science”. When I respond that I have about 20 hours of college credits in science courses, then my religion comes into play as I obviously went to a school run by “snake handlers” so naturally of course my profs were low quality or creationists. Baptist is especially guilty of the constant moving of the goal posts like that. So when mentioned my education, i wasn’t trying to play the “authority game”, which obviously you (Baptist) and Carl are. I’m all willing to continue the dialog, but lets see who throws out the first insult…
Carl states: “..On the contrary, SIU has an extremely high reputation among professional archaeologists. I do my fair share of hiring, and when I see SIU on a resume I take note…”.
I didn’t say they did or didn’t, I simply said I’d never heard of that school..I’ve heard about Clemson, USC, Ga tech, Iowa State and Bob Jones U, but never SIU..sorry about that…
Baptists states: “…I’m no expert on the Clovis issue. But I did learn about it in numerous archaeology classes. Like Carl Feagins and others have discussed here, my memory of how it was discussed by archaeologists does not mesh with Graham Hancock’s portrayal of it…”.
No one here I’m guessing thinks you are an expert on the Clovis First Model; did you actually read Hancock’s analysis of it, if so, how did he portray it?
Baptist states: “…because I’m not going to waste any more time angering and confusing you with facts and logic…”.
Project much? You are the one here who is so obviously angry, if not, why all the insults? Are you an SJW or something similar?
Baptist: Apparently, at that unaccredited school you attended in Flyover Country, “Southern Illinois University”, there were no requirements to take a course in logic nor to even read anything you disagree with, so I’ll give you a brief summary on how Hancock “portrayed” the Clovis issue in America Before; Hancock was not so much interested in the details of the Clovis Culture, because obviously those details are so well known and attested to, why bother? Rather, he was more interested in the thoughts of those who vehemently defended the Clovis Fist/Only Model, and why. Secondly, he was also interested in what came before the Clovis culture, and why that data was missed for nearly a century. Do you see the distinction? I have the book in front of me, so feel free to ask questions. But be honest, did you even read a single word that Hancock wrote, or are just relying on what his critics say here? Since you are a known liar, I’ll wager that you’ll lie about this also. “Cheers”..
Carl,
Go Salukis!!! Your insights into SIU compared to others nicely illustrates the knowledge gap here. I guess that some people might find it worth wading thru about 50 mini-tantrums to learn that SIU is a fine school for archaeology especially if one is interested in CRM. Glass half-full, I guess. I’ve lost touch with people there. I assume that they still do the annual conference sponsored by the CAI. Have you ever been to one?
I haven’t as yet, but our conference opportunities in the Forest Service are limited. There’s some belief in Washington that an academic conference is something akin to a convention at Las Vegas instead of where professionals can gather, share methods, successes/failures, and learn from each other about modern standards of the field. I’m hoping to have some stuff to present in the way of iron industry investigations and clandestine distilleries in the near future, however. Possibly even some work related to vernacular gravemarkers in cemeteries (we have over 270 historic cemeteries on my Forest) and early 20th century silage practices. I get more and more caught up in historic versus prehistoric archaeology these days!
Baptist states” “…However well your education, introductory anthropology course from a bible college..”.
You are a special kind of stupid; as I made clear, my intro to Anthropology course was taken at The College of Charleston; do you think that is a “Bible College”? Unreal…
Baptist, “headed for the tall grass..”…
Baptist states: “…I’m quite confident that I have forgotten more about just about anything discussed here than you ever learned in your entire college career…”.
Oh yeah? Well my Dad can beat of your dad! Isn’t that about the level of discourse you aspire to? But anyway, as I made clear here, I majored in US history and minored in Secondary Education; so why in the world would you want to pick a fight with me on the subject of the “1870 Federal population census”?
Further, is that tied to “anything that is discussed here”? You sound like a little uptight and light in the loafers. For what it’s worth, my specialty area within US history would be the American Civil War, and within that, the subject of Ironclads. But it’s nearly impossible to imagine, that on an archaeology blog, I would challenge you to a duel in that rather narrow field. Anyway, I have America Before right in front of me, and I’m assuming you have your copy, so if you would like to get back to that, I be more than happy for a civil dialog with you, with Carl as a slightly biased referee. With your phantom Phd from “SIU University” (is that like Phoenix University, an on-line school?). it should be a walk-over…..
Since Happily Lapsed Baptist has been completely transparent about his education and published papers, I would like to do the same amount mine, and the readers can judge which is more impressive:
1.) Attended West Hills Jr. College, Coalinga, Ca, 1986; while there I used scale models of the Bismarck and the HMS Hood to recreate their famous clash in WWII; voted “super cool” by my roommate.
2.) Attended Chapman College in 1987, presented a class presentation on the Monitor vs. Merrimack naval battle; voted “really cool” by same roommate.
3.) Attended College of the Sequois in 1989 and gave lecture entitled “Bigfoot: What the Establishment is Hiding”; voted Best paper by Visalia, Ca., “Bigfoot Association”
4.) 2008, Completed course work for “UFO Field Investigator” by MUFON (Mutual UFO Network).
5.) 2009, graduated with degree in “Ancient Technology” from the David Hatcher Childress Foundation.
6.) 2012, “Why UFOS are Real”, published in the peer-reveiwed journal “Atlantis Rising”.
7.) 2017: won an on-line debate with The Friendly Atheist.
8.) 2018; finished reading Graham Hancock’s “Supernatural”, but nearly died of a drug overdose doing it.
I think those “credentials” stack up very nicely with old Lapsed Baptist…
“QS World University Rankings by Subject 2016 – Archaeology”
Trying to find more info on this “Southern Illinois University” and it’s alleged reputation; in the above ranking of the 100 best schools of archaeology worldwide, SIU didn’t make the list. In fact, I can’t find them on ANY list of highly rated schools in that area…
“Bachelor’s in Archaeology: Top 20 Values”
Nor did SIU make this list, but guess who did? You guessed it, that “Bible School”, The College of Charleston………so it seems like, someone here is either ignorant or just a flat liar….
Carl states: “..On the contrary, SIU has an extremely high reputation among professional archaeologists. I do my fair share of hiring, and when I see SIU on a resume I take note…”.
If that’s true, Carl, I’m having a very difficult time finding that info; surely they would have made someone’s top 20/50/100 list, correct? Call me a skeptic, but can you provide me with any info outside of you just saying it, that would support what you’re suggesting?
A couple comments, not based on all these prior comments but on the review. 1. mtDNA is never considered to be “human DNA” in genetics and biology because it does not contain any human DNA. It has DNA to duplicate itself only, just like our intestinal bacteria has DNA to duplicate itself. It is simple. It is pretty clear now why you don’t understand this. 2. We don’t write to please or be accepted by the mainstream community. If we did, no one but your small closed community would read it. And it would be amazingly boring, repeat what all of you are indoctrinated to believe, and perpetrate the lies spewed by the mainstream. So understand that in the community for which we write, having the mainstream attack–especially using lies and distortions that can be easily demonstrated–is a badge of honor that demonstrates we are on a good track. For your own reasons, you assume others would want your approval. 3. Last, mainstream American archaeology rests on a bedrock of racism that began at the start of “archaeology” and the Smithsonian’s involvement. This racist base permeates throughout the entire mainstream belief system. There is a psychological process at work there I don’t expect you to understand, but as you peer out of your belief system you see racism as the explanation of people who disagree with your underlying beliefs. Maybe you should write that book you have been talking about.
All I see is a lot of projection. But then, I’m sure you would casually say the same in my direction.
There is no “mainstream” in modern archaeology. One is either doing archaeology scientifically or one is not.
Your circular reasoning here might be correct. Maybe mtDNA is never considered to be human DNA. Except by just about everyone who actually works with mtDNA professionally. For example:
Chang DD, Clayton DA (1985). Priming of human mitochondrial DNA replication occurs at the light-strand promoter. Proceedings of the National Academy of Science, USA 82, 351-355
Payam A. Gammage , Lindsey Van Haute , and Michal Minczuk (2016). “Engineered mtZFNs for Manipulation of Human Mitochondrial DNA Heteroplasmy.”In Mitochondrial DNA: Methods and Protocols, Matthew McKenzie (ed). New York: Humana Press, p. 145.
Kasiviswanathan R, Gustafson MA, Copeland WC, Meyer JN (2012). Human mitochondrial DNA polymerase gamma exhibits potential for bypass and mutagenesis at UV-induced cyclobutane thymine dimers. Journal of Biological Chemistry, 287, 9222-9229.
Laurent Chatre , Benjamin Montagne , and Miria Ricchetti (2016). “A Single-Cell Resolution Imaging Protocol of Mitochondrial DNA Dynamics in Physiopathology, mTRIP, Which Also Evaluates Sublethal Cytotoxicity.” In Mitochondrial DNA: Methods and Protocols, Matthew McKenzie (ed). New York: Humana Press, page 50 (fig.1).
Miyako K, Takamatsu C, Umeda S, Tajiri T, Furuichi M, Nakabeppu Y, Sekiguchi M, Hamasaki N, Takeshige K, Kang D (2000). Accumulation of adenine DNA glycosylase-sensitive sites in human mitochondrial DNA. Journal of Biological Chemistry, 275, 12326–12330
Spelbrink JN, Li FY, Tiranti V, Nikali K, Yuan QP, Tariq M, Wanrooij S, Garrido N, Comi G, Morandi L, Santoro L, Toscano A, Fabrizi GM, Somer H, Croxen R, Beeson D, Poulton J, Suomalainen A, Jacobs HT, Zeviani M, Larsson C (2001). Human mitochondrial DNA deletions associated with mutations in the gene encoding Twinkle, a phage T7 gene 4-like protein localized in mitochondria. Nature Genetics 28, 223–231
…and the list could go on and on, obviously.
But don’t take my word for it:
“Human mitochondrial DNA (mtDNA) is a 16.5 kb, circular, multi-copy genome encoding several structural components of the respiratory chain, ATP synthase, and RNA components necessary for mitochondrial gene expression” (Gammage, Van Haute, and Minczuk 2016).
The mainstream archaeologists here, and their supporters, who have hurled charges of racism at those who disagree with them (and that includes me), are certainly not going to like it at all when they are accused of the same sin. My guess is, in their defense, they will not only categorically deny it, but go so far to say that such a thing among themselves or in their work is utterly preposterous. In fact, it’s unthinkable. Why as we all know, racism, in all it’s forms, is something that exists on “the other side”, but certainly not here. I am going over to Amazon in a few minutes and buy the book by Little and Collins for sure….
Carl states: “…There is no “mainstream” in modern archaeology. One is either doing archaeology scientifically or one is not…”.
Dr. Albert Goodyear, chief archaeologist at the Topper Site, with whom I think you are in disagreement with, mainly because some of the artifacts he uncovered might date to around 50,000 years BP; is Dr. Goodyear “mainstream”, or in fact a pseudo-archaeologist? Given your either/or philosophy, he has to be one or the other, as their is, by your logic, no third position….
Every reference you use says “Human Mitochondrial DNA” meaning the mitochondria in our cells as opposed to the mitochondria in every other living organism. Following your reasoning they should just say “human DNA.” “But don’t take my word for it: “Human mitochondrial DNA (mtDNA) is a 16.5 kb, circular, multi-copy genome encoding several structural components of the respiratory chain, ATP synthase, and RNA components necessary for mitochondrial gene expression” (Gammage, Van Haute, and Minczuk 2016). Gee, why does it say “mitochondrial gene expression” and not human gene expression? Maybe because mtDNA produces mitochondria and not human cells? You are not worth any more of my time. On the other hand, I assume that here you are writing and asserting official positions for the US Forestry Service and the “Land Between the Lakes” section as that is prominently displayed here as well as on your Facebook and Twitter pages. No where do I see that the statements are solely your own.
Honestly, I can’t imagine I was ever worth your time. I’m just some guy on the internet with an opinion, which, by the way, is mentioned on my About page as being my own and not that of my employers’ or affiliated universities. LOL. And “taking note” is hardly a “hiring decision” by itself.
Anyway, I’ve no real dog in the hunt regarding mtDNA’s status as either human or non-human. I poked at it a bit because I was curious why it was important to you. I’m assuming there might be some grand, pseudoscientific conclusion that rests on it being this way, but damned if I can sort it out. Maybe not… you just seem unusually hung up on it. And me, for that matter. Anyway, Cheers, Greg.
Plus, on this page you discuss how you make hiring decisions for the US Forestry Service.
Carl,
I understand the situation. I’ve met people in similar straits who had to deal with a lot red tape and skepticism just to attend a conference two states away. By the way, historical sidenote my first introduction to this pseudo archaeology crap came in one of Jon Muller’s archaeology class. He didnt use that concept to describe it though. If my memory serves me correctly Muller had done some work in kentucky on glyphs or something like that. Or maybe it was Gumerman. Hard to keep track given the number of well known people who were around back then.
Baptist states: “…:Hard to keep track given the number of well known people who were around back then…”.
Well, you know, Baptist, when you make stuff up as you go along, it would indeed be difficult to keep track of all those fantasies in one’s head, like your phantom college degrees, or the assertion that the College of Charleston is a “Bible School”, but hey, on the internet you can say, or be, whatever you wish, and most people won’t ever notice the difference….
My, my, Gleaner63 has racked up quite the obsessive, oops, I mean impressive number of posts here. A few comments before leaving Gleaner to whatever he is struggling to do here.
1.I provided the links to articles demonstrating that there are debates about the nature and properties of electricity. Go back and revisit the original discussion of this, then move the goalpost back where it belongs, and then think it through. No need for me to discuss the implications of these debates for how electrical engineers work in the present because it isn’t relevant to the original discussion. Now, how electrical engineers will have to deal with the changing state of knowledge in the future is a different and more relevant topic. I guess when some things believed by electrical engineers turn out to be wrong 50 or 100 years from now then it means they are pseudoscientists. Darn, where have I heard this argument before?
2.Don’t know why you keep trying to associate me with abusing drugs. I do research on habitual substance abusers like Hancock. You can’t seem to grasp the difference.
3.I know of SIU-C by reputation and interacting with people from there (see below). A solid program with good faculty that provides four-field training and a large varied curriculum. The Center for Archaeological Investigation is well regarded. Doesn’t matter if the department is ranked 1st or 150th it still trumps a no-name Jesus school in education in anthropology or anything else. No need to even waste time going into the merits of LSU or University of Illinois compared to…I can’t even remember the names, sorry. Once you get to the Ph.D. level nobody really gives much thought to where the undergraduate was earned anyway, so only someone like you would try to make an issue of this.
4.As the following write-ups help to illustrate, Hancock does indeed misrepresent Pre-Clovis/Clovis First to try to paint archaeology as a whole in broad strokes. This is very evident throughout social media where Grammy and his advocates consistently speak in general negative terms about archaeology, or whatever discipline can be mined for a strawman argument. You’ve done this as well in other discussions here, but are now trying to change your tune for obvious reasons. Hancock has to do this because all archaeologists, at least the ones who are even aware that he exists, think Hancock is full of shit no matter where they stood on Clovis First or anything else. In fact, one of the few things that involve a legitimate consensus that tops Clovis First by a long shot in archaeology is that Hancock is a loon and a crank.. So he and his followers have to speak in generalities about archaeology although as usual they sometimes start to wiggle and claim “that’s not what he/I really meant” when the BS flag is thrown. But then a week later they are back to spouting the same old stuff. The Hancock reddit page is full of this. Kind of like intellectuallly feeble wack-a-moles. Gleaner63 is a lost cause but hopefully others with a bit better grasp of the obvious will find this interesting.
http://www.jasoncolavito.com/blog/graham-hancock-to-archaeologists-you-guys-are-the-pseudoscientists
https://www.andywhiteanthropology.com/blog/is-graham-hancocks-new-book-really-as-much-of-a-yawner-as-it-sounds-like-it-is
Lapsed Baptist,
We may have some overlap at LSU. Does the nickname Rambo ring a bell when it comes to archaeologists? How about the Library? Is the person that you mentioned at Clemson named Bob Coggeshall? I know him slightly. Ph.D. from SIU-C whom Clemson hired and granted tenure. Damn, come to think of it, Andy White at University of South Carolina has an M.A. from SIU-C. Guess that the people in South Calinky whose opinions count the most in such matters think of lot of what the department at SIU-C produces.
Gleaner63 takes the hissy fit up to an entirely new level in 3-2-1……
James Ford states: “1.I provided the links to articles demonstrating that there are debates about the nature and properties of electricity…”.
Then you must being having a debate with yourself, because I never raised the issue of the ultimate nature of electricity, and I made the point that EEs on the job don’t either. I now have about 30 years worth of experience with EEs, and for all the stuff I’ve helped them troubleshoot and repair, not once has the subject of the “nature and properties of electricity” ever arisen. So again, where this is coming from only you can tell us…
James Ford states: “…2.Don’t know why you keep trying to associate me with abusing drugs. I do research on habitual substance abusers like Hancock. You can’t seem to grasp the difference…’.
You raised the point about your alleged “drug research”, again for reasons unknown to me, on an archaeology blog. Since you continually do it, and the two have no known connection, I poke a little fun at you. What now, you can’t take a joke or 2? And what is your proof on Hancock’s drug abuse?
James Ford states: “..3.I know of SIU-C by reputation and interacting with people from there (see below). A solid program with good faculty that provides four-field training and a large varied curriculum. The Center for Archaeological Investigation is well regarded. Doesn’t matter if the department is ranked 1st or 150th it still trumps a no-name Jesus school in education in anthropology or anything else. No need to even waste time going into the merits of LSU or University of Illinois compared to…I can’t even remember the names, sorry. Once you get to the Ph.D. level nobody really gives much thought to where the undergraduate was earned anyway, so only someone like you would try to make an issue of this…”.
So you freely admit then, that SIU is not ranked by anyone as an outstanding school, and is only known by you, and that’s what matters, correct? Come on James, you can do better than that…as you said about where I graduated from, SIU is a “no-name school”…sorry…
James Ford…..well, you did get one thing correct, Clemson is highly ranked and highly regarded; but doesn’t it being in the Deep South and attended by a lot of conservative Christians somehow disqualify it? Hissy fit? Wow, do they have mirrors at your institution, or is the bong smoke just a little thick in there?
James Ford states: “..Hancock has to do this because all archaeologists, at least the ones who are even aware that he exists, think Hancock is full of shit no matter where they stood on Clovis First or anything else…”.
You’re having a hissy fit: I expected more of a reasoned argument from such a highly educated person as yourself, James….
So what really eats away at James Ford, Carl, Lapsed Baptist and others here so much that they have to accuse others or racism, drug abuse and anything else they can possibly imagine? They can’t bear the thought, even for a moment, of being wrong, and by a wide margin, on topics like the Clovis First/Only model. And that Hancock reminds them of it pokes them in the eye with it…..
Imagine belonging to a discipline, where, for the better part of 100 years, despite the best efforts of thousands or field workers, professors, professional societies, peer reviewed papers, academic journals, unlimited funds and the support of numerous colleges and universities, that at the end of the day you would have been proven to be utterly…..wrong. And not just by a few years, but tens of thousands of years. And yet this accurately describes the discipline of archeology and it’s relation to the Clovis First Model. And no doubt accounts for most if not all of the vitriol directed at Hancock and others…
James Ford states: “..Once you get to the Ph.D. level nobody really gives much thought to where the undergraduate was earned anyway, so only someone like you would try to make an issue of this…”.
I never made an issue of where anyone went to school because I didn’t see the relevance of it, but your fellow traveler, Lapsed Baptist, apparently does, along with you. So once again, you hurl insults at people, then throw a fit when someone returns the favor….
James Ford states: “…a no-name Jesus school…”
Ya just gotta love that bit of “acceptable prejudice” in the religious bigotry department, perfectly ok with you and your friends here, the kind that lecture others on racism and with such a straight face…
James Ford states: “..My, my, Gleaner63 has racked up quite the obsessive, oops, I mean impressive number of posts here…”
Your last screed James, took up almost 50 lines of text here; a little obsessive maybe on your part? Naaaa, couldn’t be. Since your posts are usually rambling, incoherent, and off topic, it’s easier to deal with them in sections, hence what you see as obsessive….at any rate, back to your crack pipe and another imaginary “research paper on drug abuse”….lol…
James Ford states: “…Grammy and his advocates consistently speak in general negative terms about archaeology..”.
Ah, I see, the tribalism thing, archaeology cannot be criticized, that would be “off-limits”; and yet Hancock is the leader of a “cult” according to you. Good grief, continued use of mary-ju-wana over a life time really does have side effects…
James recommends: “http://www.jasoncolavito.com/blog/graham-hancock-to-archaeologists-you-guys-are-the-pseudoscientists
https://www.andywhiteanthropology.com/blog/is-graham-hancocks-new-book-really-as-much-of-a-yawner-as-it-sounds-like-it-is“.
Never heard of them. Where did they go to school? What religion are they members of? Are they in a cult? What section of the country are they from? Are they now, or have they ever been racists, voted for racists, or closet racists and do they read racist literature? Unless you can answer that to my satisfaction, I can only assume they have something wrong with them. Sorry, but I’m just trying to apply your standards to who or what is or isn’t acceptable to be read or discussed here…
James:
My B.A. from SIU and some nice letters of reference from Handler, Muller, and Barton (history) got me a full ride scholarship for the LSU grad program. I’m happy with it. As I indicated before with some very good people in the right positions it doesn’t need to be an elite program to work out very well. Coggeshall was finishing up his diss. and working as a lecturer or visiting prof, can’t remember which, when I was there. He was one of the first people I met on a campus visit. He had just gotten the position at Clemson, a secular school by the way, for the following year when I got there. At least I think that is the chronology. Oh yes, Rambo. I got along better with her than a lot of people. She is hell on wheels in oral comps but I assume that you already know that unless you were geography side. I was there for Kniffen-West Fest if that helps with time frame. I had a few at the Library but was more of a Chimes person. We probably shouldn’t get too much more detailed than that in a forum like this. You can just imagine what filled my email in-box for a week the last time I was dumb enough to provide too much info in a setting like this. too many people with unhealthy fixations.
Hancock has a certain rat-like cunning and sometimes even he can figure out that it is time to leave a particular sinking ship like he did with the earth crust displacement and ancient white civilizations spiels that he was pushing a while back. I’ll bet a Dixie beer that he beats the Clovis drum less harder once he has made his bundle off the latest book. At least a lot of his fans besides the zealots are inclined to back off of it when pushed on it. Maybe he will get the memo too.
It’s way too hard to sort through all the ravings and try to carry on conversation in this section. Perhaps we can touch bases the next time our paths cross on Carl’s blog at a point before the running of the loons really opens up.
Baptist states: “…As I indicated before with some very good people in the right positions it doesn’t need to be an elite program to work out very well…”
I think anyone who’s ever been to college would recognize that; I’m happy with my undergrad education I got at CSU in the field of history. The professor I took most of my courses under had previously taught at Princeton and the University of Virginia, both elite schools….
Baptists states: “…It’s way too hard to sort through all the ravings and try to carry on conversation in this section…”.
Carrying on a civil conversation should be easy, here or anywhere else. Of course it helps to adhere to certain rules, like not purposely insulting people. It also helps to see the people you are talking to, despite the disagreements, as people to be valued, just like yourself. When Hilary Clinton referred to millions of Americans as “deplorables”, she was of course letting us know that a civil discourse was impossible, and never her actual intent anyway…
Start in with politics here, and I’ll start hitting the delete button. I detest political bickering.
..didn’t mean to make a political statement, was merely using it as a well known marker to illustrate a point…..but I’ll watch it from now on…sorry about that…
Ex-Baptist,
Dream on. Hancock’s whole civilization wiped out without a trace is just a grown man’s spin on the old elementary school “hey teacher the dog ate my homework and you can’t prove that it didn’t” routine. Clovis is just a red flag for him to wave in the face of his fans to get them so stoked up with moral outrage toward archaeology that they don’t notice that the emperor is not wearing any clothes. They then turn right around and ape the whole routine on social media. He is too invested in it, literally. Clovis ain’t going anywhere until GH heads off to that big ayahuasca patch in the sky.
I came after the big Fred Kniffen conference. Or maybe it went down when I was out of state doing research. Or maybe I am confusing it with when they hosted the Conference of Latin American Geographers or something like that. Too many moons ago. By all means let’s pick this up elsewhere and by all means no contact info online. I will be on the road for the near future so it will be more later than sooner.
Geaux Tigers!
Misunderstanding and misrepresenting genetics research are the newest additions to the pseudoscience toolkit.
James Ford states: “…Clovis ain’t going anywhere until GH heads off to that big ayahuasca patch in the sky…”.
Try to keep up, James: the Clovis First/Only Model is already gone…really, really gone. Tribal/mainstream archaeologists have now fallen back to a pre-prepared position; “…well, we’re still right, even when we are wrong..” (everything we say is “provisional”, so that should cover nicely for this and later, debacles”.
“pseudoscience toolkit.”
Mmmmm; wonder what the alleged “official science toolkit” might contain?
1.) Claim that all official positions by science are provisional, including, but not limited to: 1.) the Earth is round, 2.) Evolution is true, 3.) there is no Lost Civilization, 4.) The heliocentric solar system model, 5.) The Big Bang, 6.) The vast age of the Earth, 7.) Man landed on the moon, 8.) the Face on Mars is a natural geologic formation, 9.) ESP is false, 10.) Ghosts aren’t real, 11.) Roswell wasn’t an ET crash site, 12.) Creationism is false, 13.) Global Warming is a fact….
Try defending any of those positions as “provisional” and see what happens..
This is probably offering up pearls to swine but: there is a big difference between the process by which people from Copernicus to Sagan collect data, analyze it, interpret it, and communicate findings and how it is done by people like Collins and Little. Just because certain propositions in the past were viewed as outrageous by some it is not a valid argument for uncritically accepting every outrageous proposition that every Tom, Dick, and Harry comes along with in the present as valid or even provisional. Inundating a blog comments section with apples and oranges logic and details of underwhelming credentials only makes you look amateurish and foolish. Your sure to be multiple responses will undoubtedly be amateurish and foolish and it is clear that you have nothing better to do than to behave in this manner. So, I see no need to continue the discussion. Good evening.
Susan Ford: Thanks for the comments. Carl will probably delete this because he apparently feels the need to protect those on one side of this argument while holding those on the other side to a more rigid standard, but we’ll see. In regard to your “swine argument”, is that a good, sound, scientific argument to you? You didn’t include a picture, so I can’t judge. I’m curious however, about your comments regarding Carl Sagan. Prior to von Daniken, when Sagan believed that Paleo-Contact (or Ancient Astronauts) was a perfectly legitimate theory, what data did he “collect, analyze and interpret”? Just curious….
From Wikipedia: “…in Intelligent Life in the Universe (1966) astrophysicists Iosif Shklovsky [Shklovskii] and Carl Sagan devote a chapter to the argument that scientists and historians should seriously consider the possibility that extraterrestrial contact occurred during recorded history; however, Shklovskii and Sagan stressed that these ideas were speculative and unproven.[20] Shklovskii and Sagan argued that sub-lightspeed interstellar travel by extraterrestrial life was a certainty when considering technologies that were established or feasible in the late 1960s;[21] that repeated instances of extraterrestrial visitation to Earth were plausible;[22] and that pre-scientific narratives can offer a potentially reliable means of describing contact with aliens…”.
Apparently prior to 1968 when Chariots of the Gods was published, it was “cool” to inquire into the Paleo-Contact realm. There is no mention of the idea being pseudo-science, or that Sagan was using “pseudo-scientific methodology”; in fact I can’t find another scientist from that era denouncing Sagan as a fraud/quack/crank or maybe even a “closet racist” or possibly citing “racist sources” in his tome. Weird, isn’t it? The very pinnacle of pseudo-science, and the poster child for pseudo-science, is/was Eric von Daniken, and yet Sagan was there at the beginning. No doubt the reason for the lack of criticism was that other thing in the skeptics toolkit, that of special pleading and tribalism…oh well….
By bringing up Von Daniken compared to Sagan in this way and basing your argument only on Wikipedia as a primary source you just managed to illustrate all of Susan Ford’s points without even realizing it.
Fly on the Wall states: “…By bringing up Von Daniken compared to Sagan in this way and basing your argument only on Wikipedia as a primary source you just managed to illustrate all of Susan Ford’s points without even realizing it…”.
If there is an error in the Wikipedia article, which is based on Sagan’s book, then by all means, point out the error. Or, if you like, I can quote from the actual book itself, if that is more to your liking. Which would you prefer? If you can’t point to a single error in the Wiki article, then you are, like a lot of others on here simply blowing smoke, without even realizing it. You are either unwilling or unable to actually engage with the argument (my guess is the latter)….
Sagan made a proposition early in his career and he later modified his perspective. EVD continues to make an entire career out of slipshod research and misrepresentations that fall apart under close scrutiny. It doesn’t help his case that Von Daniken was convicted of fraud and embezzlement in relation to his career as a researcher. One could make fruit salad out of the apples and oranges logic being thrown about here.
Magnus Canis: I stopped reading EVD in the very late 70’s because I became convinced that he’d produced no actual, verifiable evidence of Paleo contact. So, as to continuing career in that field, I’ll have to take your word for it. As for his prison record, none of it has any bearing on his central theory, that’s an ad hom attack on your part. Apples and Oranges logic? Don’t keep us waiting with your thoughts……:)
Magnus Canis: A simple question that won’t require a long response from you; when Sagan, apparently in 1966, a full 2 years before EVD burst on the scene, first proposed the “Paleo-Contact” theory, was it then considered “pseudo-science”? Or maybe a “pseudo-scientific theory”?
At the time Sagan began his career, it was just becoming clear that humans could develop the technology to travel to outer space. This raised the serious question of if there were beings from elsewhere who had the same or better technology and could have visited earth in the past. Sagan took this notion in one direction based on evidence at hand and EVD took it a very different direction form day one and has continued a pattern of bias confirmation and misrepresentation for nearly 50 years now. EVD actually confessed to forging artifacts to support his case. This confession appeared on the BBC broadcast “The Case of the Ancient Astronauts” which aired on March 8, 1978. So, his criminal conduct in matters of fraud do have direct bearing on his career. If you can’t grasp the distinction then I can’t help you any more with this and neither can anyone else.
Magnus: I’m very aware of the distinction you’re making; I’m not “contesting” that issue. No one, absolutely no one, is somehow implying that Sagan and EVD, either in their training, education, career or morals are even remotely the same. For whatever reason you don’t want to address what I wrote/asked:
“…A simple question that won’t require a long response from you; when Sagan, apparently in 1966, a full 2 years before EVD burst on the scene, first proposed the “Paleo-Contact” theory, (1). was it then considered “pseudo-science”? 92). Or maybe a “pseudo-scientific theory”?…”
Do you see those questions? Neither of them have anything to do with what Sagan and EVD did *after* their theories were first proposed. If you can’t address that, there’s no reason to continue this thread
Magnus states: “…so, his criminal conduct in matters of fraud do have direct bearing on his career. If you can’t grasp the distinction then I can’t help you any more with this and neither can anyone else…”.
Again, I’m mot addressing EVD’s career, only his ideas; and no, they have absolutely no bearing on the truth of Paleo-Contact, you should be able to grasp that. As an example from history, Karl Marx was known to have, among other sins, neglected his wife and children, even to the point of abuse. In your mind, does that automatically invalidate Marxism?
Another of Carl Sagan’s Pseudo-scientific moments:
“…Sagan’s interest in UFO reports prompted him on August 3, 1952, to write a letter to U.S. Secretary of State Dean Acheson to ask how the United States would respond if flying saucers turned out to be extraterrestrial…”.
I wonder if EVD did the same? Mmmmmmm?
It looks like the comments section for an article on Denisovans and DNA is the wrong place to see a discussion of Denisovans and DNA.
Does anyone think Dean Acheson replied to good ole Carl’s inquiry? Probably just thought, “…another nut case…”…LOL…..
Lee Groff: I just bought “Denisovan Origins”; it’s sober, easily followed, and utterly unfantastic in it’s claims. I know most skeptics won’t read it, but after some of the commentary here I’m glad I purchased it to see what all the fuss is/was about (clue: you will be sorely disappointed, especially in regard to the charges of racism leveled at the authors)….